Publications
2026
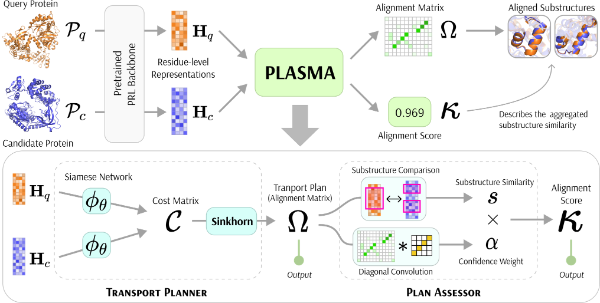
- Fast and Interpretable Protein Substructure Alignment via Optimal TransportZhiyu Wang, Bingxin Zhou, Jing Wang, and 4 more authorsIn International Conference on Learning Representations (ICLR), 2026
PLASMA is a module based on optimal transport that can be seamlessly added to any protein representation learning model with minimal training or even no training to enable fast and highly accurate identification of locally similar substructures. Extensive experiments on 7 backbone protein language models across 3 tasks from the VenusX dataset demonstrated consistent and significant improvement (+10-30% ROC-AUC) across all backbones and tasks.
@inproceedings{wang2026plasma, title = {Fast and Interpretable Protein Substructure Alignment via Optimal Transport}, author = {Wang, Zhiyu and Zhou, Bingxin and Wang, Jing and Tan, Yang and Zhao, Weishu and Li\`o, Pietro and Hong, Liang}, year = {2026}, booktitle = {International Conference on Learning Representations (ICLR)}, eprint = {2510.11752}, archiveprefix = {arXiv}, primaryclass = {q-bio.BM}, github = {ZW471/PLASMA-Protein-Local-Alignment} }
2025
- arXiv
 Topotein: Topological Deep Learning for Protein Representation Learning2025
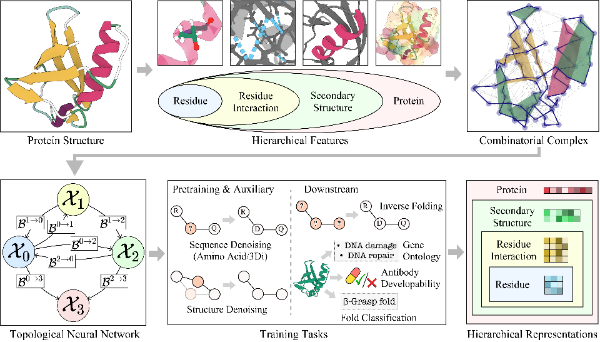
Topotein: Topological Deep Learning for Protein Representation Learning2025Topotein introduces a comprehensive framework that applies topological deep learning to protein representation learning through the novel Protein Combinatorial Complex (PCC) and Topology-Complete Perceptron Network (TCPNet). TCPNet is an SE(3)-equivariant topological neural network that simultaneously learns protein residue-level and secondary structure-level features without losing any geometric information, demonstrating consistent improvements across four protein representation learning tasks.
@article{wang2025tcpnet, title = {Topotein: Topological Deep Learning for Protein Representation Learning}, author = {Wang, Zhiyu and Jamasb, Arian R. and Hajij, Mustafa and Morehead, Alex and Braithwaite, Luke and Li\`o, Pietro}, year = {2025}, eprint = {2509.03885}, archiveprefix = {arXiv}, primaryclass = {cs.LG}, github = {ZW471/TopoteinWorkshop} } - arXiv
 MoRE-GNN: Multi-omics Data Integration with a Heterogeneous Graph AutoencoderZhiyu Wang, Sonia Koszut, Pietro Liò, and 1 more author2025
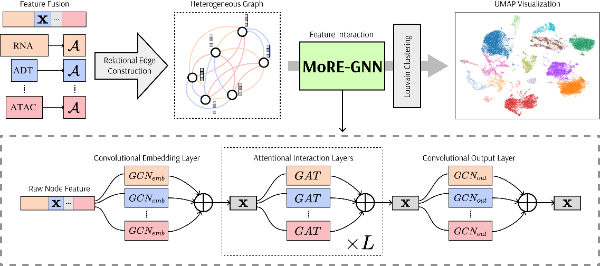
MoRE-GNN: Multi-omics Data Integration with a Heterogeneous Graph AutoencoderZhiyu Wang, Sonia Koszut, Pietro Liò, and 1 more author2025MoRE-GNN is a heterogeneous graph autoencoder that combines graph convolution and attention mechanisms to dynamically construct relational graphs directly from single-cell multi-omics data (e.g. RNA, protein, and ATAC modalities), embedding them into a shared latent space for cell clustering. Evaluations on six publicly available datasets demonstrate that MoRE-GNN captures biologically meaningful relationships and outperforms existing methods.
@article{wang2025multiomics, title = {MoRE-GNN: Multi-omics Data Integration with a Heterogeneous Graph Autoencoder}, author = {Wang, Zhiyu and Koszut, Sonia and Li\`o, Pietro and Ceccarelli, Francesco}, year = {2025}, eprint = {2510.06880}, archiveprefix = {arXiv}, primaryclass = {cs.LG}, } - GraphAU-Pain: Graph-based Action Unit Representation for Pain Intensity EstimationZhiyu Wang, Yang Liu, and Hatice GunesIn IJCAI Micro-gesture Analysis for Hidden Emotion Understanding (MiGA) Workshop, 2025
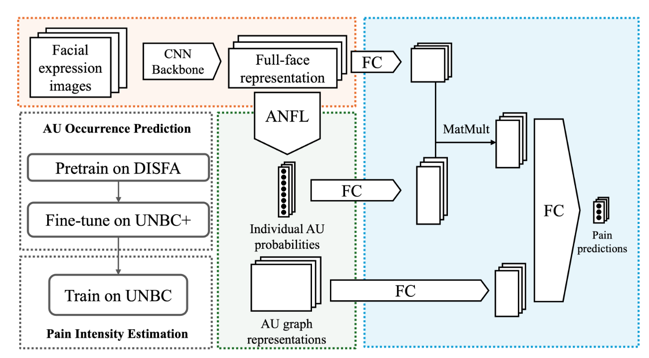
Understanding pain-related facial behaviors is essential for digital healthcare in terms of effective monitoring, assisted diagnostics, and treatment planning, particularly for patients unable to communicate verbally. Existing data-driven methods of detecting pain from facial expressions are limited due to interpretability and severity quantification. To this end, we propose GraphAU-Pain, leveraging a graph-based framework to model facial Action Units (AUs) and their interrelationships for pain intensity estimation. AUs are represented as graph nodes, with co-occurrence relationships as edges, enabling a more expressive depiction of pain-related facial behaviors. By utilizing a relational graph neural network, our framework offers improved interpretability and significant performance gains. Experiments conducted on the publicly available UNBC dataset demonstrate the effectiveness of the GraphAU-Pain, achieving an F1-score of 66.21% and accuracy of 87.61% in pain intensity estimation.
@inproceedings{wang2025graphaupaingraphbasedactionunit, title = {GraphAU-Pain: Graph-based Action Unit Representation for Pain Intensity Estimation}, author = {Wang, Zhiyu and Liu, Yang and Gunes, Hatice}, year = {2025}, booktitle = {IJCAI Micro-gesture Analysis for Hidden Emotion Understanding (MiGA) Workshop}, address = {Guangzhou, China}, eprint = {2505.19802}, archiveprefix = {arXiv}, primaryclass = {cs.LG}, }